Excitement: Possible Splice Product Found!!
Recap: What am I trying to do?
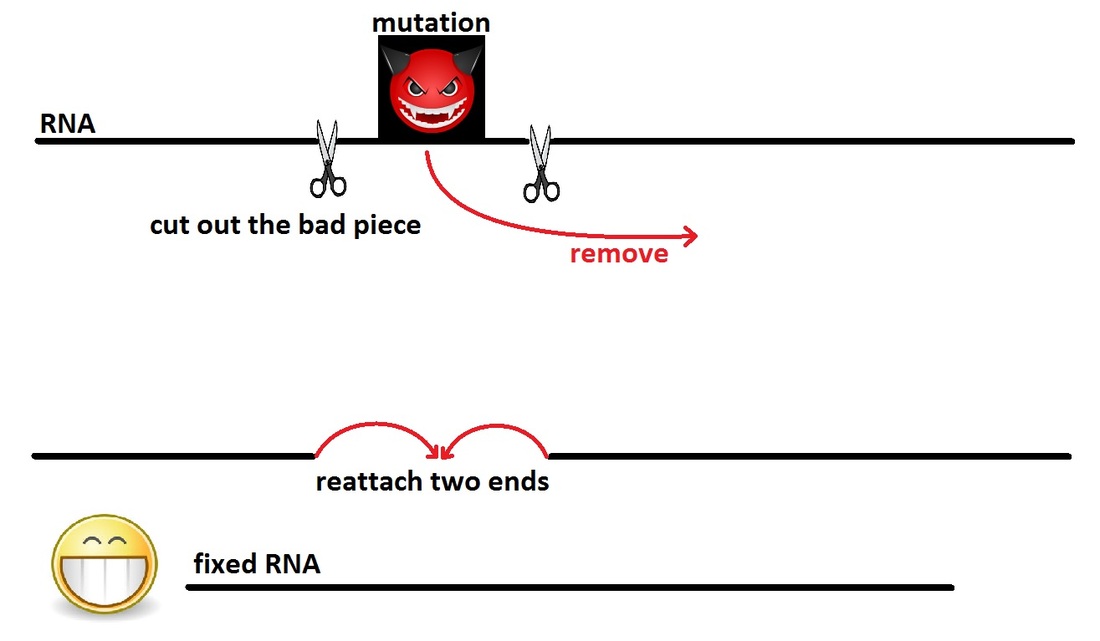
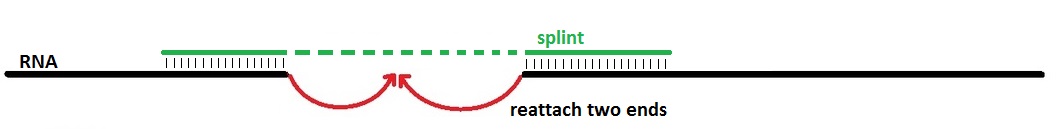
Recap: What are splints?
Back to the Story: Attempted Sequencing, FAIL!
Unfortunately, the sequencing failed due to pour quality or lack of priming. I have some ideas on what to do though.
Disappointment: Is it really my splice product?
In fact, the third gel seemed to say that I didn't have splice product. A very disappointing end. However, there might be some hope that it still is. I have to get the sequencing right to know for sure.
Soccer and Science

What a game! Tim Howard was *amazing.* At least we played hard and well, though we lost 2 to 1.
Future Plans
1) Get a clean gel of my DNAzyme digestion products.
2) Sequence my bands correctly to determine if they are splice product.
Challenges with #1
In order to get #1 to work, I have to label my RNA with a 5' iodoacetamidofluorescein (5-IAF), as I mentioned here in my last post. I need that label, because my RNA is not staining well and I cannot see it without a fluorescent tag. So far, when I've tried to do this reaction, it worked, but the RNA degraded. I'm going to try it again with an RNase inhibitor, that should block RNA degradation.
Challenges with #2
Sequencing my splice product is not easy because it is so short. It's hard to sequencing anything below 100 bp, and mine is 50 bp. Therefore, it may be necessary to insert (clone) it into a plasmid and sequence the whole plasmid. However, this route is not cheap.
The second issue is the poor quality. The Nanodrop UV-Vis spectrums of my extracted DNA were terrible. I'm not sure why they are so poor quality, since I filitered them. I'm going to try to remedy this by doing PCR on the extracted samples, cleaning up the PCR reaction and testing them again. I *should* get cleaner samples, I believe, and far more DNA. Then, I will try to sequence them one more time the regular route before I go into plasmid cloning steps.
I will be working on BOTH these issues next week.






 RSS Feed
RSS Feed
