|
Emory welcomes it's newest crop of grad students to the department! We enjoyed showing off our work with posters, talking about research, the program, hanging out and having fun! These are the third set of new students that I have seen come into Emory. It's amazing how time flies. Tonight, bowling and more!
0 Comments
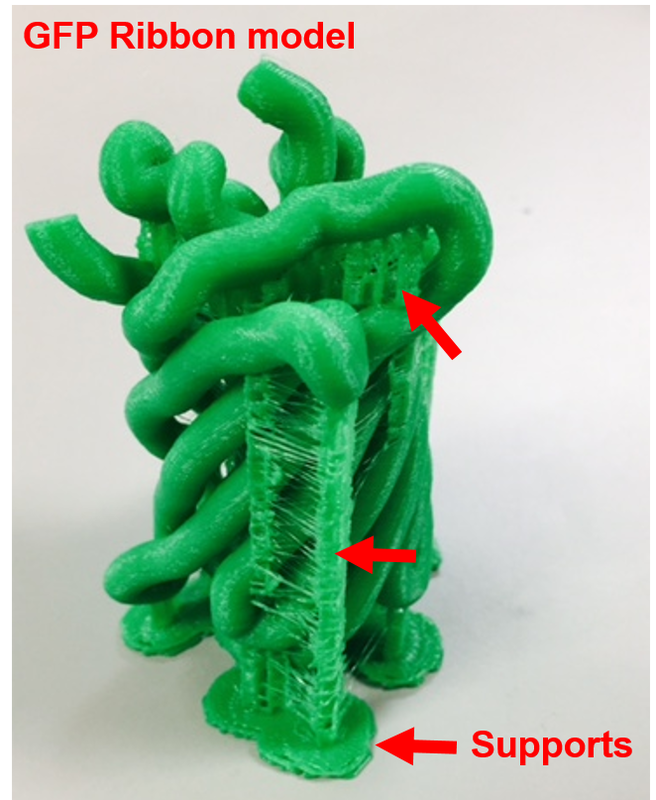
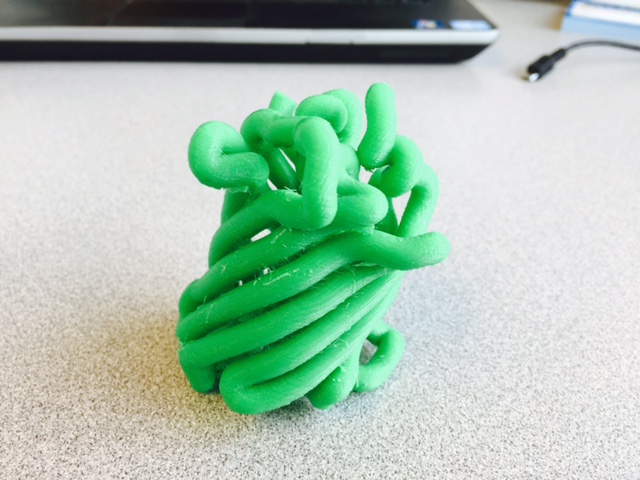
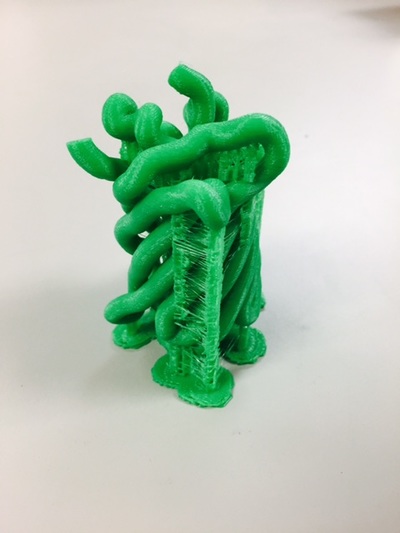
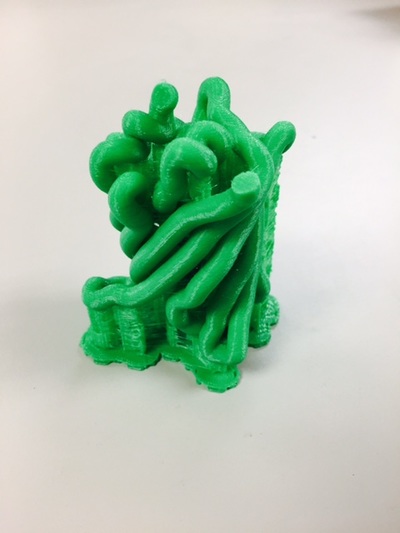
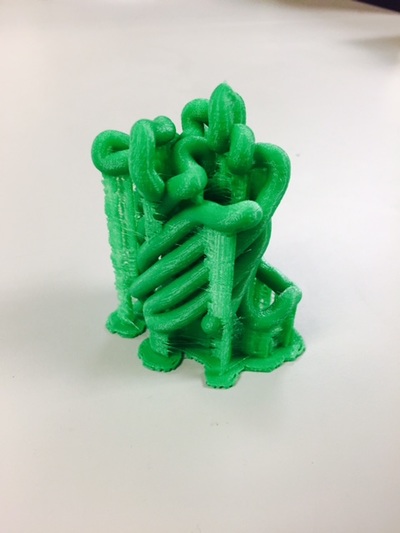
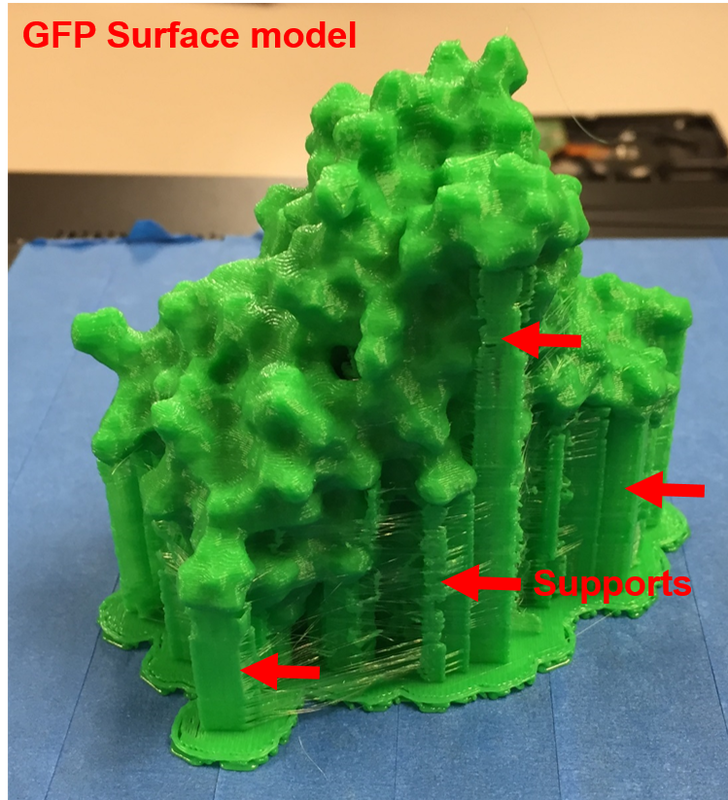
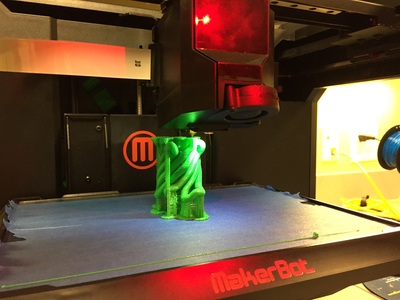
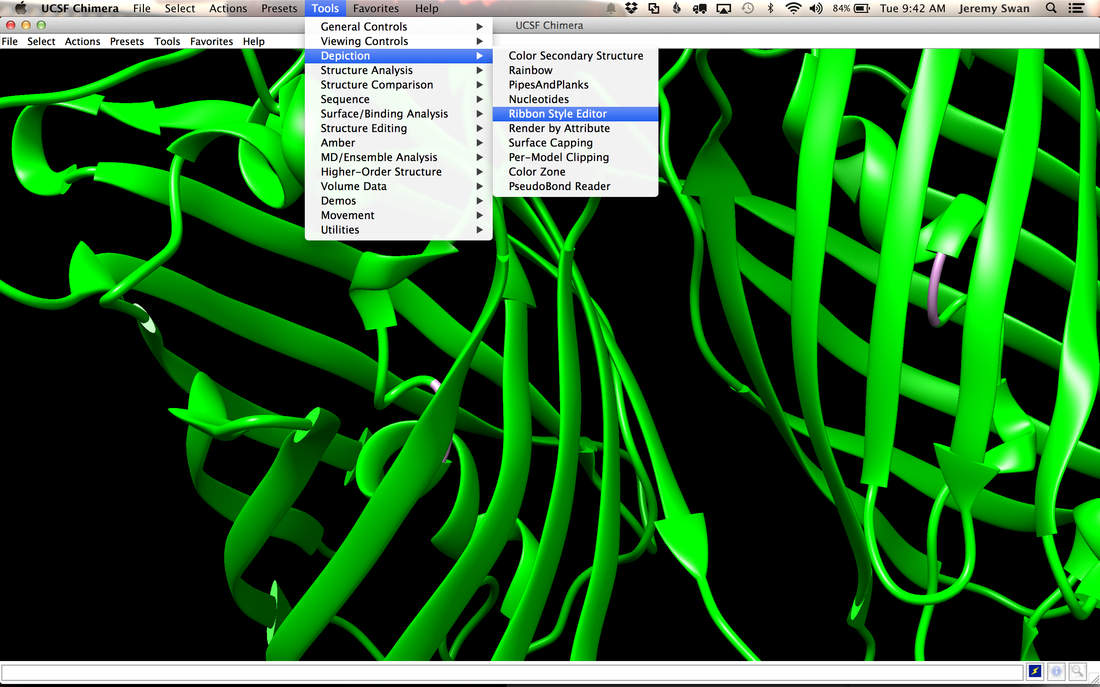
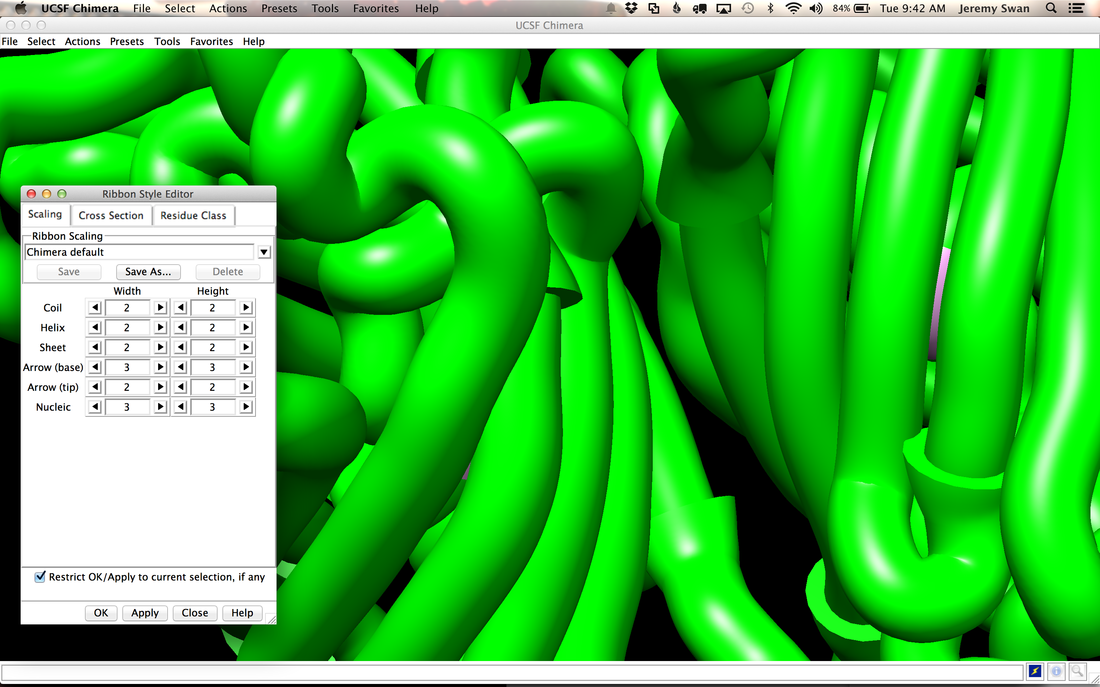
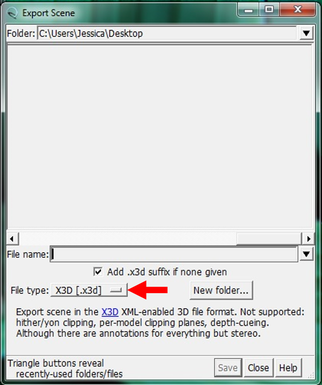
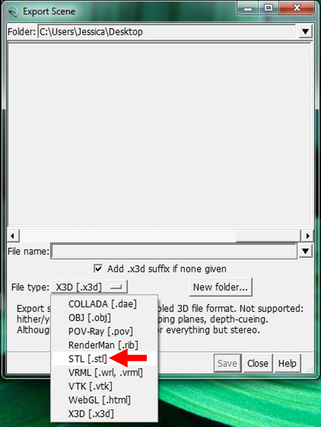
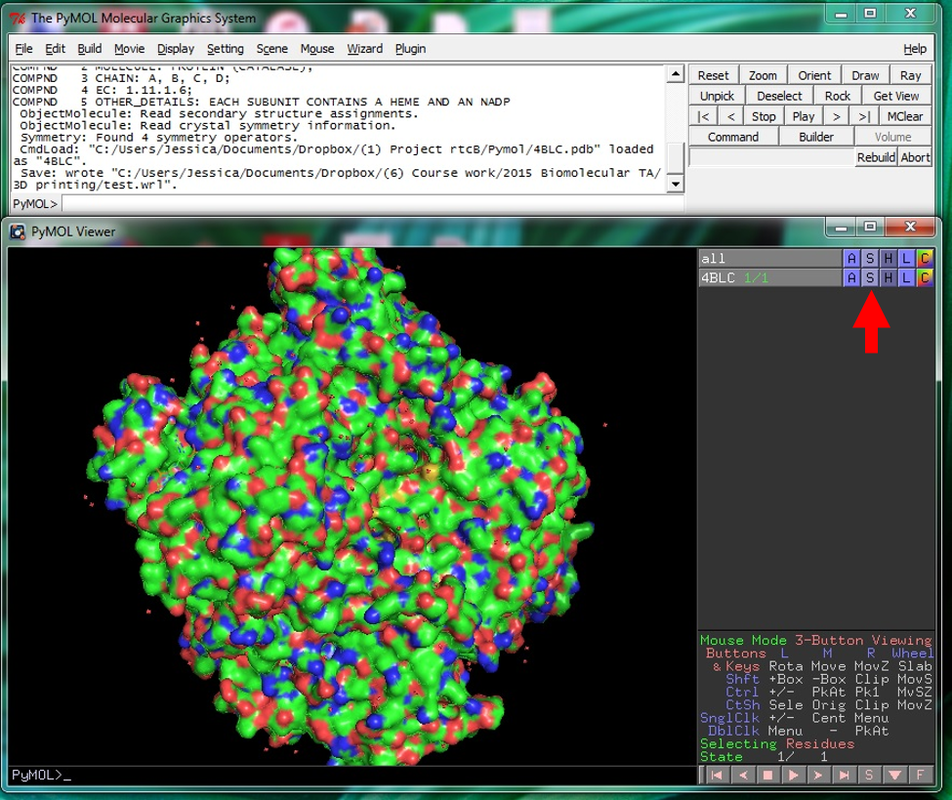
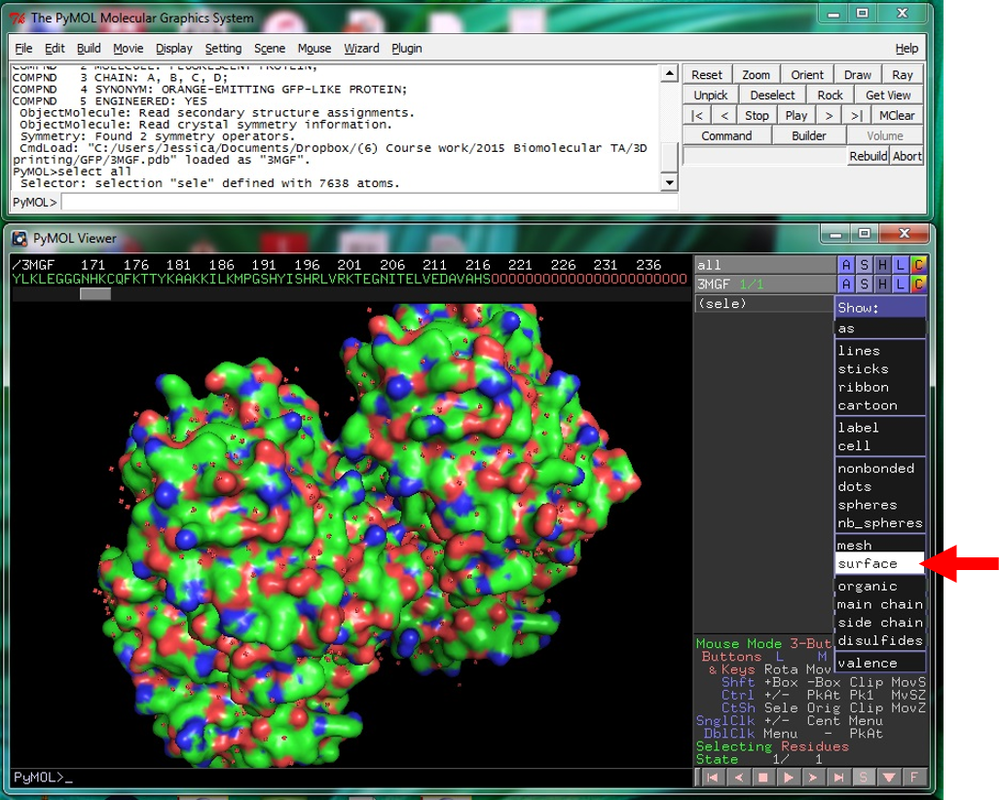
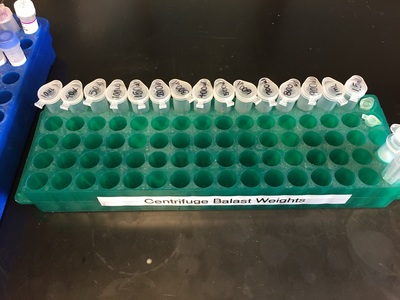
Why Mirkin is cool*from his website here 1. He is one of the founders of nanotechnology. 2. He invented dip-pen nanolithography. 3. He is is the Director of the International Institute for Nanotechnology at Northwestern. 4. He has been recognized for his accomplishments with over 90 national and international awards. 5. He is the author of over 562 manuscripts and over 930 patents worldwide (246 issued). 6. He is the founder of four companies, Nanosphere, NanoInk, AuraSense, and AuraSense Therapeutics, which are commercializing nanotechnology applications in the life science and semiconductor industries. 5. He is a Member of the President’s Council of Advisors on Science & Technology (PCAST, Obama Administration). 6. He is the only chemist (one of only 15 scientists, engineers, and medical doctors) to be elected to all three US National Academies (the Inst. of Medicine, the Natl. Academy of Sci., and the Natl. Academy of Engin.). Photos! Kornelia shows off her modelVictor's presentationKornelia's presentationYun's presentationAfter presentations are overMirkin's presentationThis post has been long in coming. On July 31st, our whole lab went ziplining from 2 pm until 6 pm! Four hours! We went through difficulty levels 1 - 4 of historic banning mills - the longest zipline tour in the world! The level 4 was intense. I am deathly afraid of heights and it was only with the constant love and support of my labmates that I was able to make it through all this. I'm glad I did it. I feel like I conquerored a giant. Enjoy the photos! My (*Special thanks to Swanjere who outlines how to thicken ribbon models for printing here. I've copied and linked his advice in the following post.) You can 3D Print Your Own Protein! Here's One Way HowDETAILED INSTRUCTIONS 3D PRINT SHOP at EMORY UNIVERSITY Location: 2nd floor, Cox Hall, the computer tech lab Directions: Go up the stairs to Cox Hall, to the floor above the cafeteria. There will be a sign for the computer / tech lab. Upon entering the second floor, it’s the first door on your left. After entering the center, go all the way to the back past the desks / computers and take a right. You will run into the 3D print shop. Printers: Anyone can print objects on one of the five 3D printers available: 4 PLA plastic printers (two Taz, a Makerbot and one other) and 1 resin printer. PLA is a biodegradable plastic. Resin can be printed in finer detail than PLA. All you need is a .stl file. I’ll go over where to get those in the next sections. Printing an object and cleaning up the model: When printing an object, the printer prints supports underneath parts of the object that are in mid-air. These supports must be broken off and removed after printing with clippers / tweezers. The staff in the print-shop will help you orient your protein to have minimal supports. Some supports may be printed in the interior of the object, if it is a hollow ribbon structure. Try to remove as many of these as you can. I recommending printing on the PLA printers, as removing resin supports is more challenging. For a video of the GFP ribbon model being printed, see this video here. Each PLA print costs $2.00, regardless of size, and each resin print costs $4.00. CHOOSING A PROTEIN: SOURCES Chose a protein structure that you want to 3D print from one of the following sources: a. NIH 3D Print Exchange here b. Thing-i-verse, 3D print library, here c. Protein Databank (PDB) here Using one of these sources, chose a protein structure – not a small molecule – try to choose a protein that is not gigantic and is a monomer. The NIH 3D print exchange and Thing-i-verse give downloads of their objects directly as .stl files, however, their database is limited. If you are interested in a protein that is not in these databases, you can acquire it from the Protein Databank, which has the structures of all known proteins. The Protein Databank structures are downloadable as .pdb files, which can be converted into .stl files. Specific directions from each of these sources is given below. CHOOSING A PROTEIN: TYPE OF PRINT, SURFACE OR RIBBONS It is up to you whether you want to choose a surface model or a ribbon model to print – depending on what aspect of the protein you want to highlight and whether or not that particular aspect is visible in either kind of model. Both surface and ribbon models can be downloaded from the NIH and Thing-i-verse. However, if you want to do ribbon models, see extra note on ribbons below. Note on Ribbon Models: Ribbon models are more difficult to print, as they are more delicate. In order to print a ribbon model, I recommend using the ribbons format of the GFP model that I printed from the NIH database here. However, there is only one of these types of ribbons on NIH. The other types of ribbons available I have not tested. However, there is a method to convert any .pdb file into the thicker GFP ribbons, if you want to do that, which I’ve listed below. SOURCE 1-2: NIH 3D PRINT EXCHANGE and THING-I-VERSE From the home page of the NIH 3D print exchange, hit “Discover” and you’ll get to their library of 3D structures or click the link given, which takes you to this page. Click on a model you’re interested in. Hitting “download” under the photo will give you a wide range of options. Download the .stl format of the type of model you want. One can search Thing-i-verse and download objects in .stl format in a similar manner. Just click the item, and “download this thing!” to get your .stl file. SOURCE 3: PROTEIN DATABANK, HOW TO CONVERT .PDB file into .STL (RIBBONS / SURFACE) Downloading from the Protein Databank The majority of you may need to choose protein from the Protein Databank. For best success, chose smaller proteins under 80 kDa. After finding your protein of interest, click the title, click “download files” and download PDB File (Text). NOTE: Make sure that you download the structure of a protein monomer. Often times, when these proteins are crystallized, they are crystallized as two or three proteins together. Such structures are more difficult to build models out of. Finding a proper monomer structure may take some trial and error. Converting your .pdb file into ribbons that can be printed in .STL format You’ll need to download UCSF Chimera here. Open your .pdb file. The thicknesses (and heights) of ribbon geometry components can be easily adjusted in Chimera by clicking Tools > Depiction > Ribbon Style Editor. Making the "height" and the arrows, which interconnect the various parts thicker and more robust, will ensure it's easier to print and won't fall apart when removing support structures (see screenshots). Saving your file as .stl: After you’ve converted to the thick ribbons shown in this presets in the above screenshot, save your file by clicking “File” --> “Export Scene” --> click the box next to file type (red arrow) --> and chose “.STL” from the list (red arrow, see screenshots below). SURFACE MODELS: Converting your .pdb file into a surface model To convert your .pdb file into a surface model .stl you’ll need to download 2 programs: Pymol and Meshlab. Pymol: Download the .pdb file from the Protein Databank (listed above). Download the program Pymol here. Download “from students and educators” and fill out the forms for download. Once downloaded, open your .pdb file. Meshlab: Meshlab can be downloaded from the site here for either Mac or Windows. Getting a surface model: Open Pymol. Open your .pdb file. In the right-hand window, click the “S” (show) and click “surface.” Wait while it loads the surface format. It takes a few minutes. Afterward, click File --> Save Image As --> VRML 2 … Open the VRML 2 file in Meshlab by going into File --> Import Mesh. Save the file as a .stl by hitting File --> Export Mesh As --> hit the drop down menu and chose .STL. You now have a .STL surface model! PRINTING THE MODEL AT THE PRINT SHOP On the day of your time slot, arrive at the print shop (see direction above) with your .STL file on a flash drive. They will set you up on a printer, have you sign a waiver so you can use the tools to trim the supports after printing, and have you pay a $2.00 print fee on your Eagle Card for PLA prints. Make sure you have money on your Eagle Card! Money can be added to it on OPUS. Scale: Request the 3D printing staff to scale your model to be 8-10 cm on one side, either tall or wide, so that it is a decent size to be handled. Pick up your protein the following day and trim the supports. If the print fails, you will not be charged. GFP Ribbon Model!I printed this GFP from the NIH 3D printing database, downloading the .stl file here, and it turned out great! The original maker of this print details there the process I describe for making the thickened ribbons for a more robust ribbon model. If you Google search 3D printing proteins, you'll also come up with many other handy guides. Mine originates and pulls from many of these. Happy printing! GFP Surface Model!Fun moments from July! It's been a while since I've given an update! So here are some of crazy things that have happened this month, chronicled in pictures. LEFT: Photo of my RNA stocks in the -80C freezer. This is what they look like! I store them in tiny amounts here. This month, part of my work has included continuing to organize the -80C freezer stocks and update our list of growing plasmids, thanks to Kornelia's hard work and her student Mara. Research Lately: Mired in difficulties after big successesI got my first publishable data in March / April and it was VERY exciting! However, since then, I've been unable to make more RNA due to it degrading - some of my reagents have gotten contaminated - and I'm trying to sniff out the problem in the coming week! Very important. I'm also helping Grace to put together a poster of her work. Grace working on a reaction to transcribe RNAOrganizing the ballastsThe centrifuges need ballasts - so Grace and I worked on organizing them and making sure that they were accurate! I synthesized a third batch of gold nanoparticles!People in the Conticello lab draw creativelyPlaying with dry ice!Enzymes that come in the mail sometimes come with dry ice! Here's an example of us playing with some. I observed flip-flop cookies and I can't remember where fromThe Weinert lab uses an excess amount of Ni:NTA beadsJim Vickers in the Hill Lab graduated
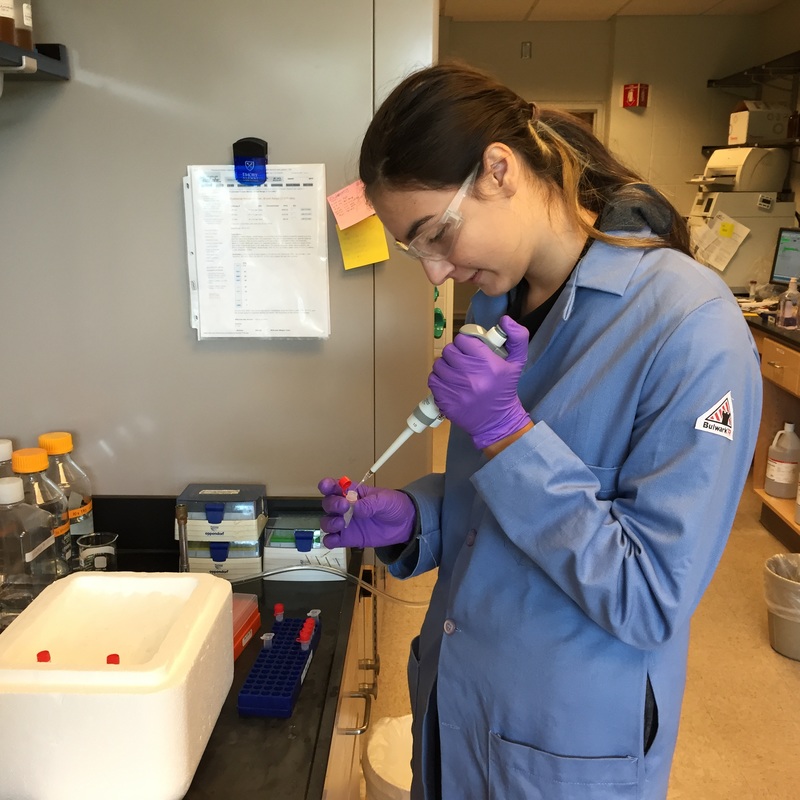
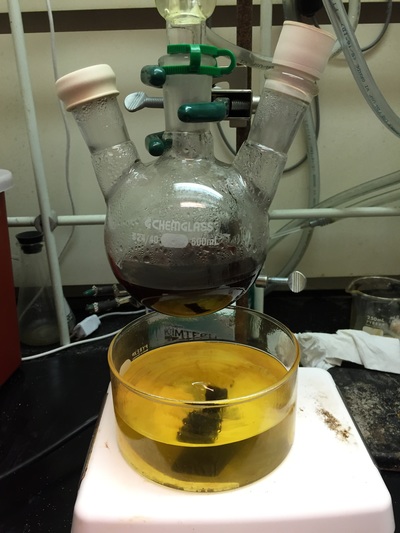
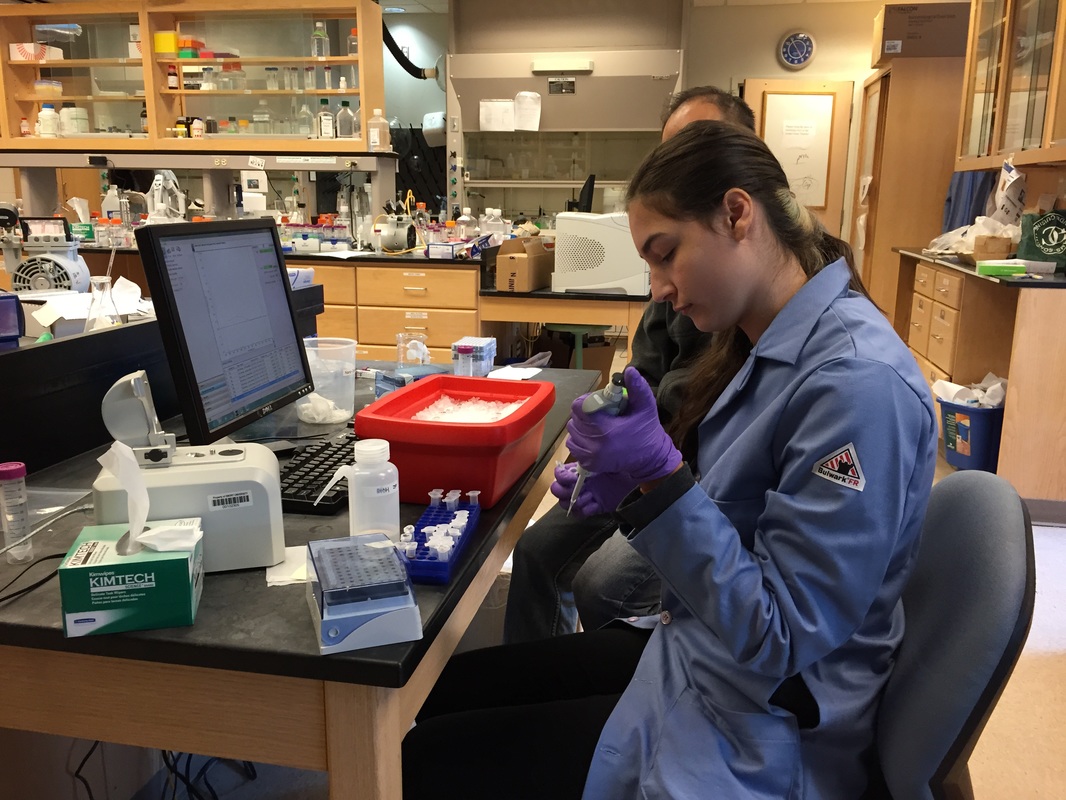
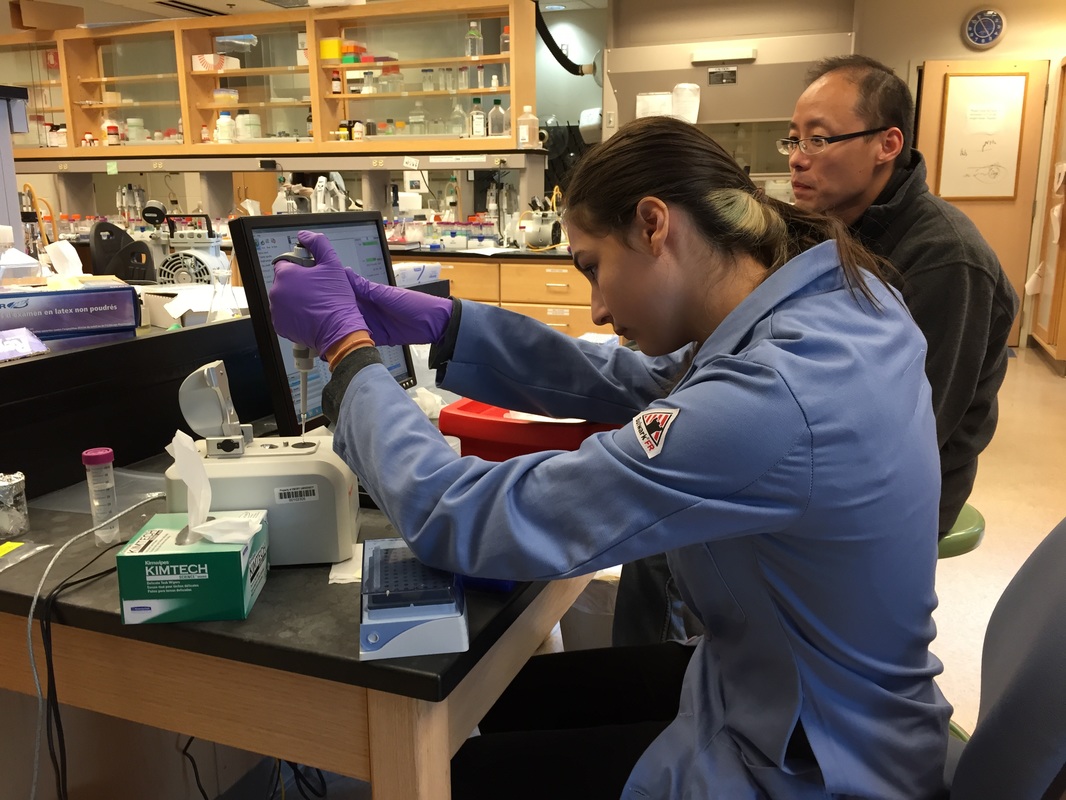
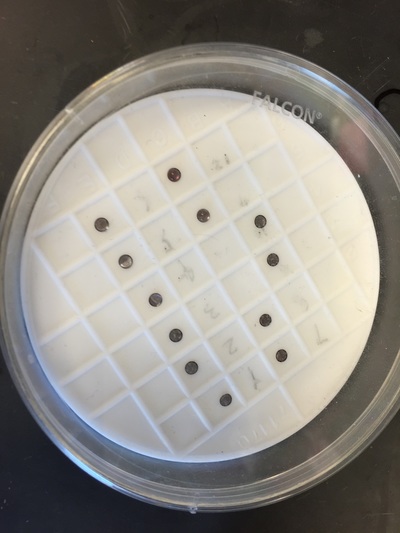
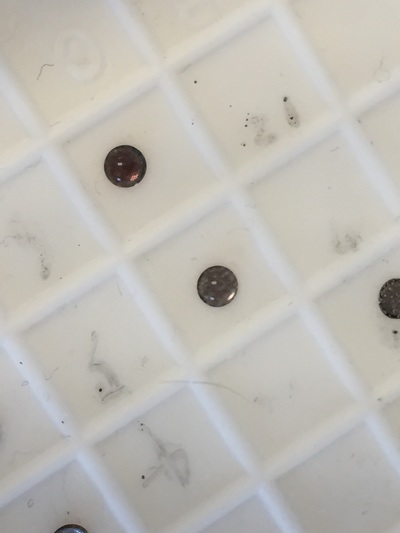
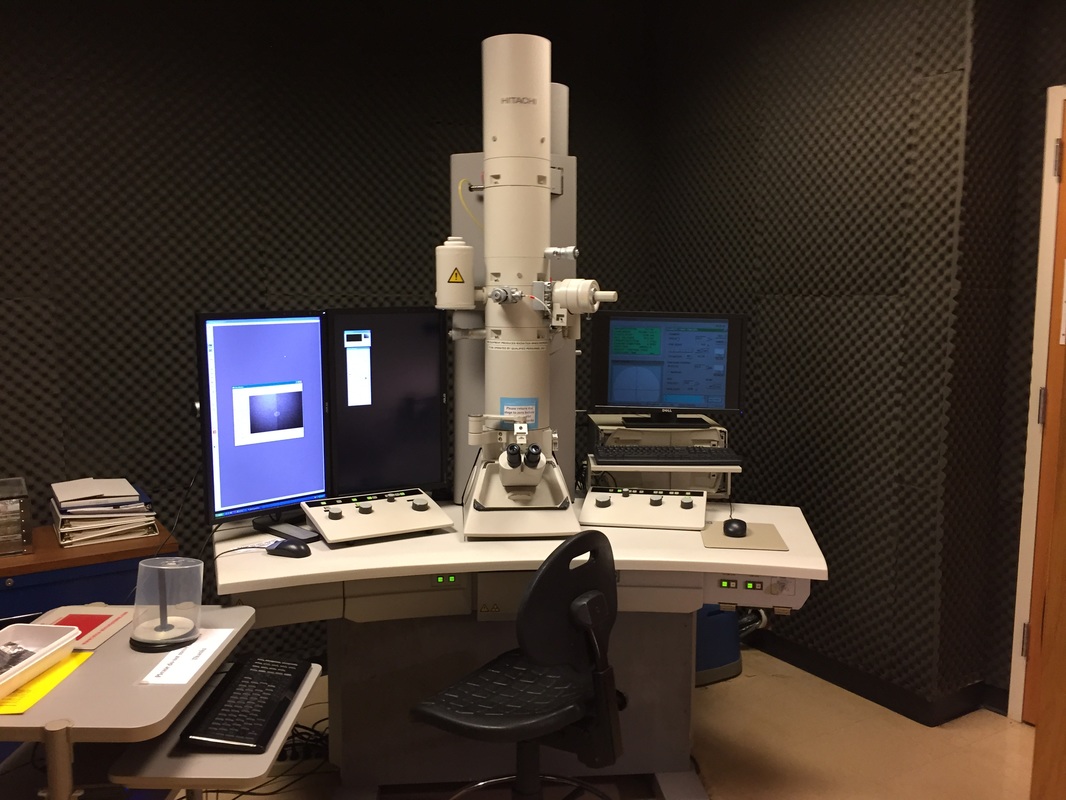
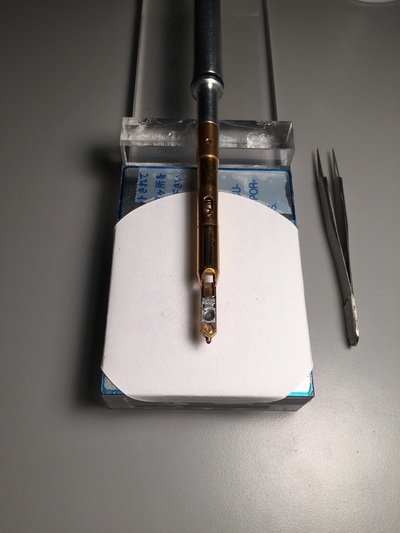
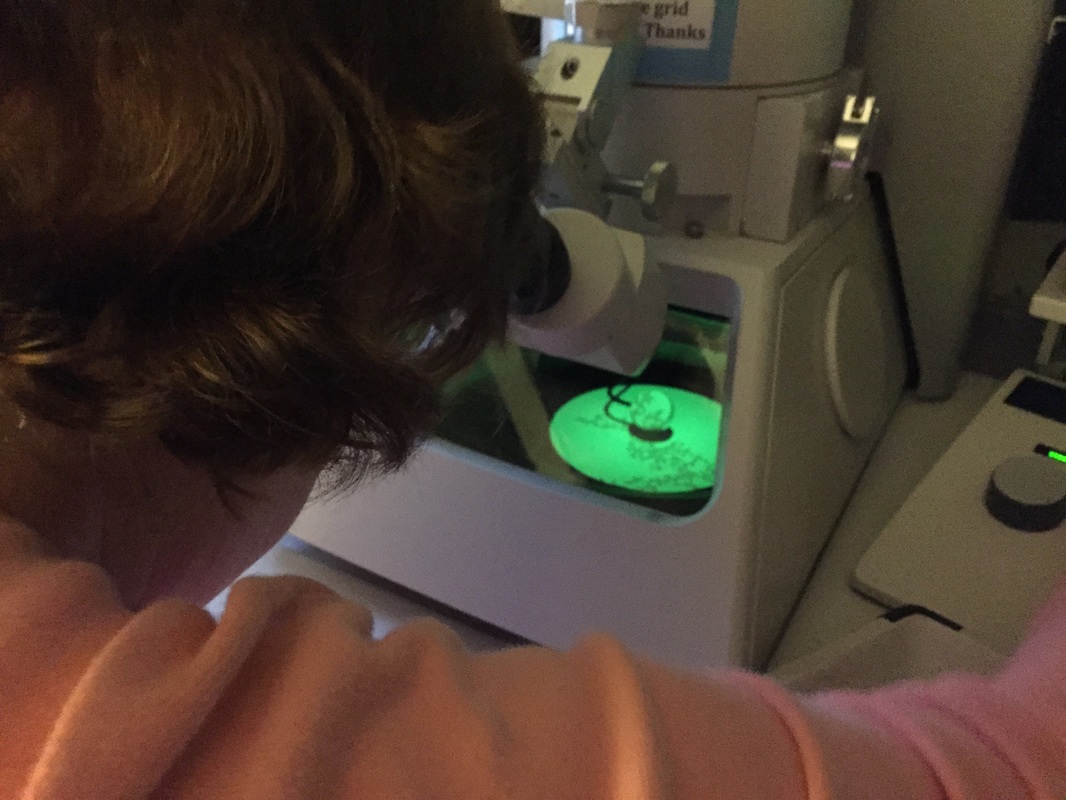
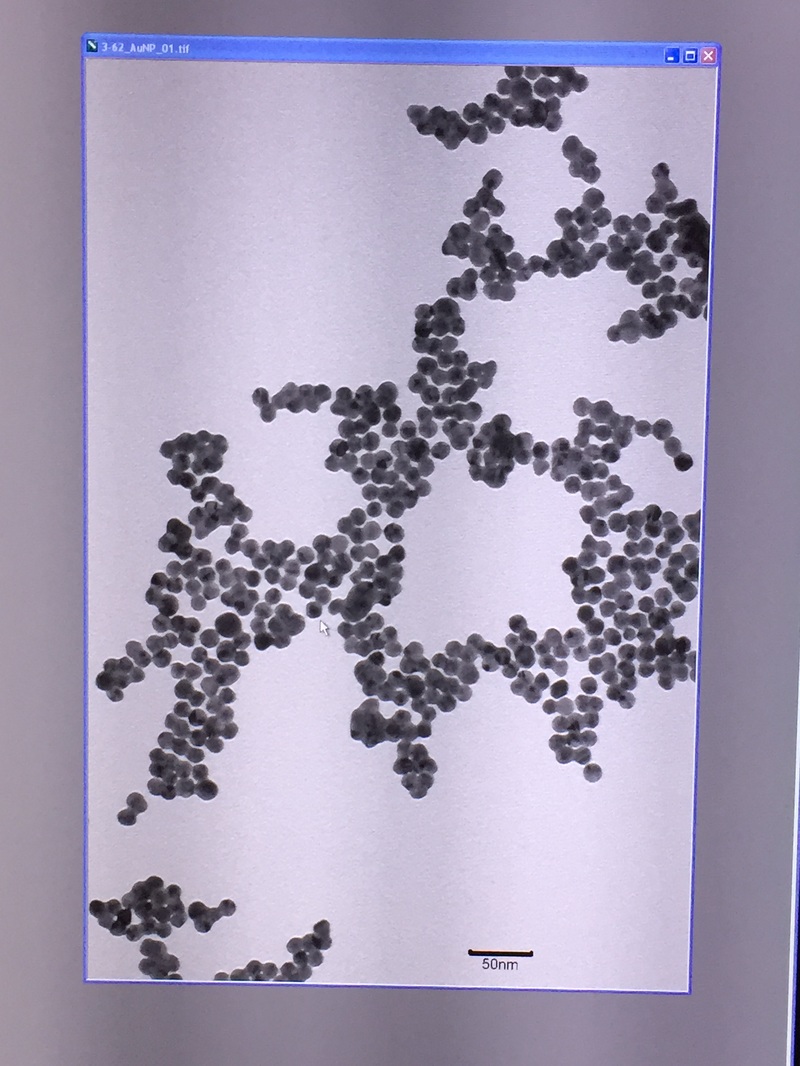
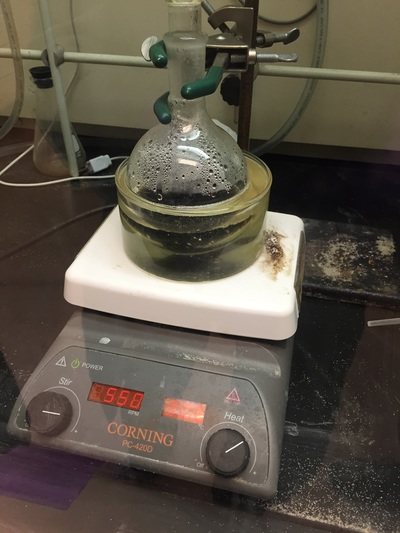
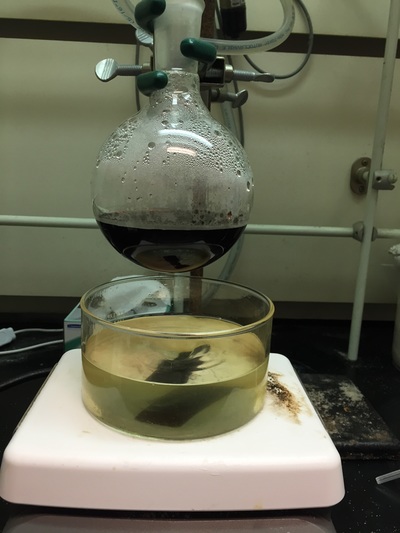
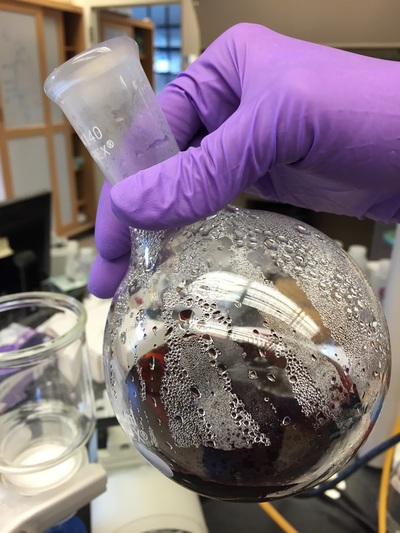
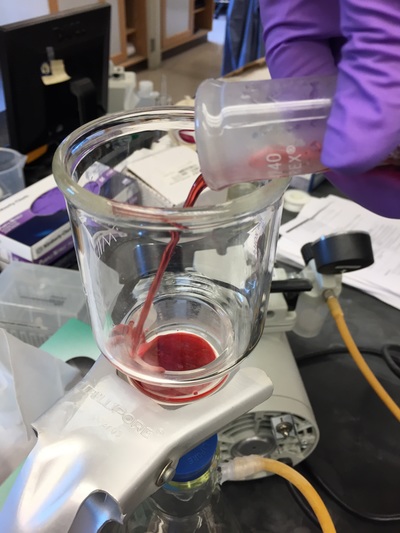
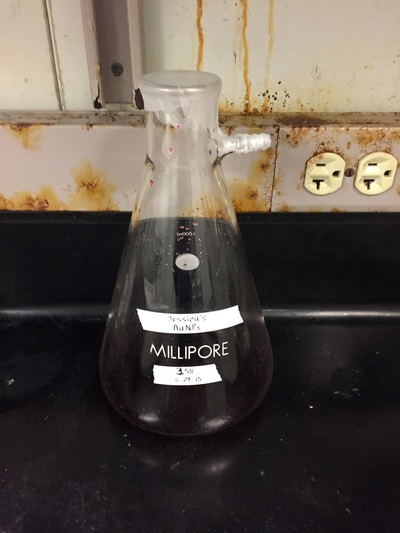
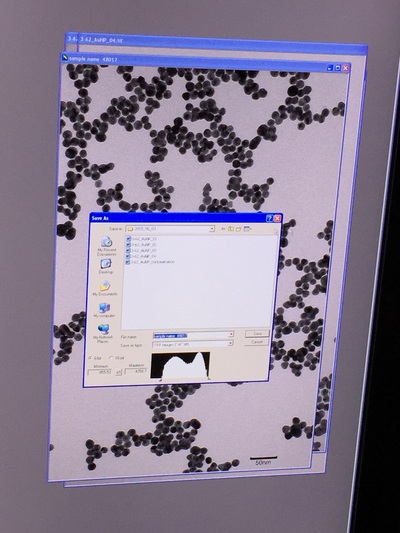
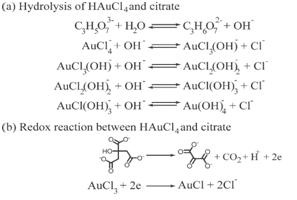
The Defense Kevin's defense was amazing -- I loved it! His data was really interesting, and he showed additional data I didn't even know that he had with his nanorollers. The party afterward was great too -- I had way too much chocolate covered marshmallows and made myself slightly sick. What's up now? Kevin is currently looking for a post-doc position, and right after his defense left for a conference to mingle amongst the scientific community and see what's out there. He's not quite leaving yet, remaining with us until he gets a post. That's great for me, because he can teach / guide me on how to do some of the techniques in his papers -- a kind of passing of the torch, if you will. My project is a spin-off idea of Kevin's that he was never able to pursue, so discussions with him have been integral to my work. Photos of the Defense and Party AfterwardGrace Thomsen joins me for the summer! I have the pleasure of welcoming Grace to the Salaita Lab! She will be working with me this summer in our lab until August, as part of my home school mentorship program, and is helping me characterize the enzyme I'm using in my work. This week was her first! Previously, she came once a week to shadow me, and finally became official upon taking Emory's safety training seminar. This week, we've enjoyed expressing my enzyme in E.coli: preparing the buffers for the expression, doing the expression, and today isolation and purification of the resulting protein. Our post-doc Weiwei is also learning protein expression to get his feet wet in some biochemistry. Previously, he almost exclusively worked with quantum dots, nanotech and inorganic chemistry. Grace also got to witness the synthesis of gold nanoparticles! We've been having lots of fun. I look forward to a fun and carefree summer full of SCIENCE! More updates will be coming soon. Here, Grace measures the concentration of protein via NanodropPreparing the sampleDrop 5 ul of AuNPs onto appropriate grid (ours is carbon). Wick away excess liquid after a few minutes with a wedge of filter paper (not a kimwipe, like what I did). Imaging the sample!I did not do the imaging myself, as I've not yet been trained on the transmission electron microscope. This scope looks at things smaller than a wavelength of light (like my nanoparticles) by bombarding them with electrons. Placing the grid in the sample compartmentTaking imagesConclusion: Some of my particles are fused, so I will repeat the synthesis this Tuesday -- oh well -- more practice!!Kevin Yehl teaches me his mad skillz for particle making before graduating!*Protocol comes from the 2006 Mirkin paper, "The bio-barcode assay for detection of protein and nucleic acid targets using DTT-induced ligand exchange." Among other things, it demonstrates how to reduce gold salt with sodium citrate to produce ~13 nm gold nanoparticles. This post chronicles my FIRST ATTEMPT at making Kevin's 13 nanometer gold nanoparticles (AuNPs)!! I've been wanting to do it for two years -- and I was SO EXCITED!! I am going to repeat the procedure next Tuesday to see if I can get them to be a little more pure. By TEM, it seems some of these are fused. Glassware and Materials NeededGlassware / Materials 500 ml flask (for gold salt solution) 100 ml flask or beaker (to hold sodium citrate soln before it's needed) condenser filter funnel 500 ml flask that attached to filter funnel 0.45 um acetate filter oil bath magnetic stir / hot plate oval stir bar (for gold salt soln) paper clip / stir bar (for oil) Chemicals aqua regia hydrogen tetrochloroaurate(III) trihydrate sodium citrate Nanopure water Step #1. Rinse the glassware used for making the particles with aqua regia to remove all trace metals -- VERY DANGEROUSThis is the step that's no fun. Aqua regia is code for "highly dangerous, concentrated and corrosive acid." Quote from Wikipedia: "The mixture is formed by freshly mixing concentrated nitric acid and hydrochloric acid, optimally in a volume ratio of 1:3. It was named so because it can dissolve the noble metals gold and platinum." Kevin showed me the glassware I needed to rinse with this stuff, and then I did it. It took me exactly 1 day -- to think about it, read about aqua regia, buy extra gloves, plan how I was going to do it, and then do the actually deed, which took about maybe 1 hour. Step #2. Rinse glassware in copious amount of Nanopure water.This is to remove the acid with which you just rinsed it. Step #3. Prepare a solution of hydrogen tetrochloroaurate (III) trihydrate.Also known as gold(III) chloride trihydrate (see here) -- your gold salt -- this is where the gold comes from. Prepare 500 ml of a 1 mM solution. I did a 1/5th recipe of 100 ml of this solution. It's a happy yellow color. Do NOT use a metal spatula when preparing any of the reagents used in this recipe. Step #4. Boil the gold salt solution in an oil bath until refluxing at 1 drip / sec.Make sure that both the solution itself is stirring with a stir bar, and the oil bath is stirring (in this case, with a paper clip in it), to prevent bumping. Ensure that the condensor is on and flowing. Go ahead and turn up the heat to 300C, otherwise it takes forever. You want to get it to reflux vigorously (boil so it condenses and drips back down) -- 1 drip / sec. See here for a video of it stirring. Step #5. Prepare the sodium citrateDo not use a metal spatula! You need 50 ml of 38.8 mM sodium citrate tribasic dihydrate if you're using the original 500 ml recipe. Step #6. Add the sodium citrate once reflux is achievedThis is where the fun starts! The solution will turn from yellow to clear, to black to purple to deep red as the sodium citrate reduces the gold into 13 nm nanoparticles. After 15 min, turn off heat and allow reaction to cool to room temp (2-4 hrs). Filter the solution in the all-glass filter / holder with 0.45 um acetate filter. See this video for a filtering attempt -- before I got the vacuum pump to work. Transfer gold nanoparticle solution to clean storage bottle. Store at room temperature or at 4C.
|
AuthorWhat's going on in my corner of the science world? Find out here! Archives
March 2019
About CategoriesLinksCategories
All
Rolling Statuses: Technical journal blog. Here you may discover what the daily life of a grad student looks like: day-to-day snippets of life, clutter, rolling statuses and unimportant fluff.
Progress Updates: Will include entries with more meaningful science. Weekly lab report: My write-ups on what I did each week (I posted these publicly during my rotation but not as much now. That may change.) Science StatusHere is a link to collected writing, poster and presentation tips.
As of February 8, 2014 I have officially joined the Salaita lab!! Very exciting. Stay tuned for updates. "Micro Min" category equates to grad school journaling; most of these have moved to my status updates blog under Home tab. See "progress updates" on this blog for more important news.
|

































































































































































 RSS Feed
RSS Feed
