Experiments This Week
Monday – setup large ligation reaction of both the FAM and hybrid substrates comparing Shuman and Raines conditions side-by-side.
Tuesday – poured and ran an 18% TBE-Urea gel of RNA FAM ligation reaction; stained hybrid gel with SYBR gold. Imaged both on Typhoon instrument. Results seem to indicate that the Shuman conditions do affect the FAM substrate but not the hybrids. FAM substrate under Raines conditions appears active. FAM gel is fuzzier than I’d like; hybrid gel is missing lanes. I believe that’s due to running so many lanes and two gels at once (urea leaching back into the wells while loading). Proteinase K came in.
Wednesday – reading and organizing day
Thursday – recalculated all concentrations for the reactions; setup simplified FAM reaction under Raines conditions. Added RNasin. Poured and stored (4C) two RNA 10-well 18% TBE-Urea gels.
Friday – stored gels somehow acquired detachment bubbles overnight. Ran the ligation reaction with and without proteinase K treatment on one of them in spite of this. Ran a second gel (15% TBE-Urea precast, 15-well, Biorad) using formamide/EDTA stop solution I’d made in June as the loading dye. Imaged on Typhoon.
There appears to be an error – I must have added RNA 10mer2 to the “rxn, only FAM” aliquots, because product band shows up there. The samples hated the formamide dye or they were two concentrated for a 15-well gel. The RNA does not look uniform and/or slightly degraded on the gel I poured but much more clear and solidly uniform on the precast gel, therefore, I believe the degradation I am getting must come from the gel itself. Both gels indicate that rtcB is active with the FAM substrate under Raines conditions with and without proteinase K.
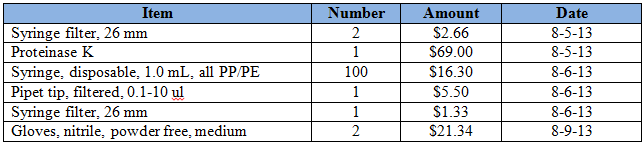
Purchases for the week = $116.13
1. The gel bubble issue is not yet completely solved. Bubbles show up after overnight storage and even occasionally small ones will appear upon immediate storage.
I will see what gels John Wang in the Gallivan lab pours – he should be doing somewhere between 15-20% and what recipe he is using. I’d like to compare it to mine and make sure that the bubbles are not resulting from some kind of improper TEMED/APS ratio. If it’s not that, it could be that our TEMED is old and is therefore not working as anticipated.
2. I am bothered that I’m getting degradation in my gels of my RNA samples. The fact that I’m not getting it in the precast gels as much indicates to me it’s a problem with my gel.
The next gels I run I’ll setup the Gallivan lab with their reagents. Perhaps we have RNase contamination in our gel reagents – Acrylamide/Bis, TEMED, APS or urea.

 RSS Feed
RSS Feed
